
deSIPHR:
de novo Protein Sequencing

The proteomics challenge
Proteomics has always faced a fundamental constraint: you can only measure what you already know to look for.
The current workhorse — mass spectrometry — requires matching protein fragments against reference databases. If a protein isn't in the database, or doesn't ionize reliably, it's invisible. Other approaches rely on fluorescent labels or antibody-based affinity methods, which introduce their own biases and blind spots.
The result is a field that has spent decades generating an increasingly detailed map of a small, well-lit corner of the proteome, while biology’s most important data layer remains hidden.
This isn't a sensitivity problem. It's a category problem. Existing tools were never designed to read proteins directly de novo. They were designed to find what researchers already suspected was there.
Pumpkinseed is built to find everything else.

How deSIPHR works
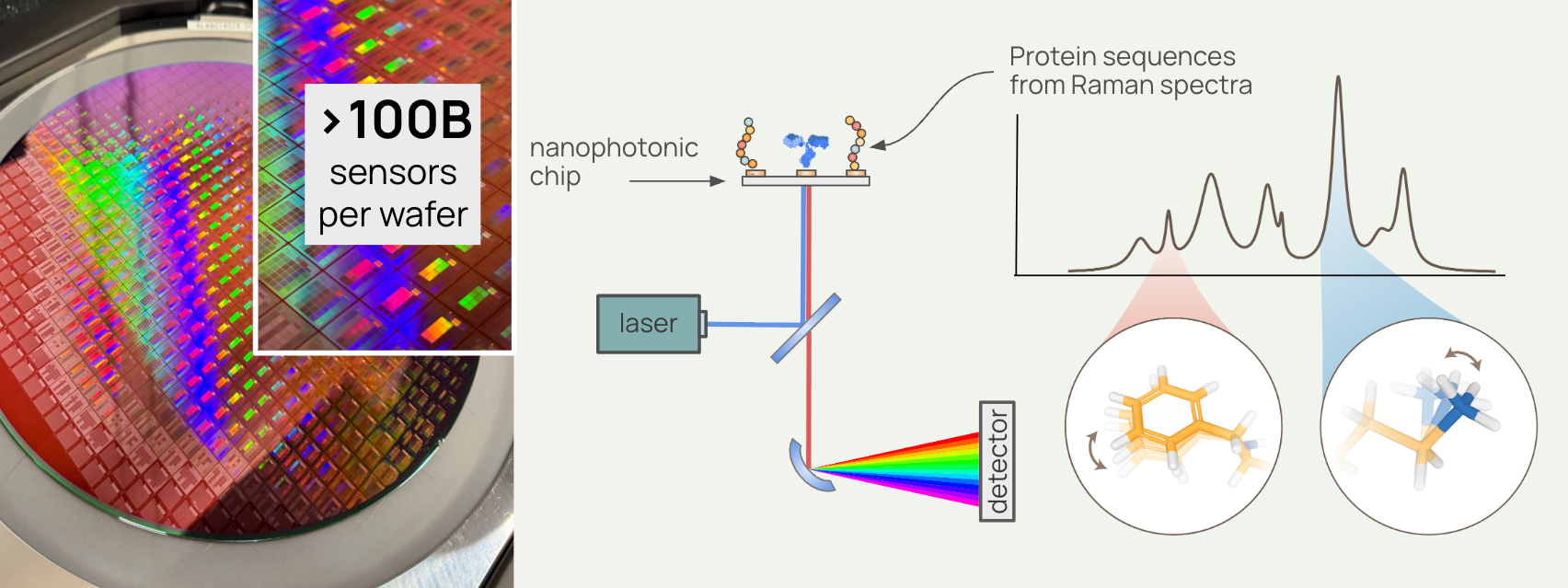
deSIPHR is Pumpkinseed's proprietary nanophotonic chip platform. With over 100 billion sensors per wafer, it reads protein sequences one amino acid at a time using Raman spectroscopy. No reference proteins. No elaborate sample prep. The result is direct, high-resolution proteomic data at a scale and fidelity that simply hasn't existed before.
What is Raman spectroscopy? Rather than tagging or fragmenting proteins, Raman spectroscopy reads the molecular vibrations of individual molecules. Each amino acid vibrates at a characteristic frequency, producing a unique physical signature that deSIPHR detects directly. This is physics reading biology in the most literal sense.
The chip itself is fabricated using semiconductor manufacturing — the same processes that produce modern microchips — which means it scales the way silicon scales. More sensors. Higher throughput. Lower cost per data point over time.



A technology built for this moment
De novo, by design. deSIPHR sequences proteins without prior knowledge of what's in the sample. No reference database required. No hypothesis about what to look for. This is the fundamental shift from confirmation to discovery.
Single-molecule resolution. The platform reads proteins at the level of individual molecules, capturing the full complexity of the proteome, including low-abundance proteins, post-translational modifications, and proteoforms that existing methods miss entirely.
Scale that matters. With over 100 billion sensors per wafer, deSIPHR generates proteomic data at a throughput that makes meaningful datasets possible. Not individual experiments, but the kind of rich, comprehensive coverage that AI biology and drug discovery actually need.
Minimal sample requirements. Meaningful results from a fraction of sample size required by traditional proteomics approaches, a critical advantage for clinical and single-cell applications.
