Predicting targeted- and immunotherapeutic response outcomes in melanoma with single-cell Raman spectroscopy and AI
Kai Chang*, Mamatha Serasanambati, Baba Ogunlade, Hsiu-Ju Hsu, James Agolia, Ariel Stiber, Jeffrey Gu, Saurabh Sharma, Jay Chadokiya, Amanda Gonçalves, Fareeha Safir, Nhat Vu, Daniel Delitto, Amanda Kirane*, Jennifer Dionne*, Predicting targeted- and immunotherapeutic response outcomes in melanoma with single-cell Raman Spectroscopy and AI, bioRXiv (2025)
Abstract
PURPOSE. Identifying reliable predictors of immunotherapeutic response in melanoma remains an outstanding challenge. Existing transcriptomic and proteomic profiling methods for the tumorimmune microenvironment (TIME) are costly and may not faithfully capture modifications actively impacting tumor behavior. Here, we present a non-destructive, single-cell approach combining Raman spectroscopy and machine learning (ML) that enables rapid cell profiling and therapeutic response prediction.
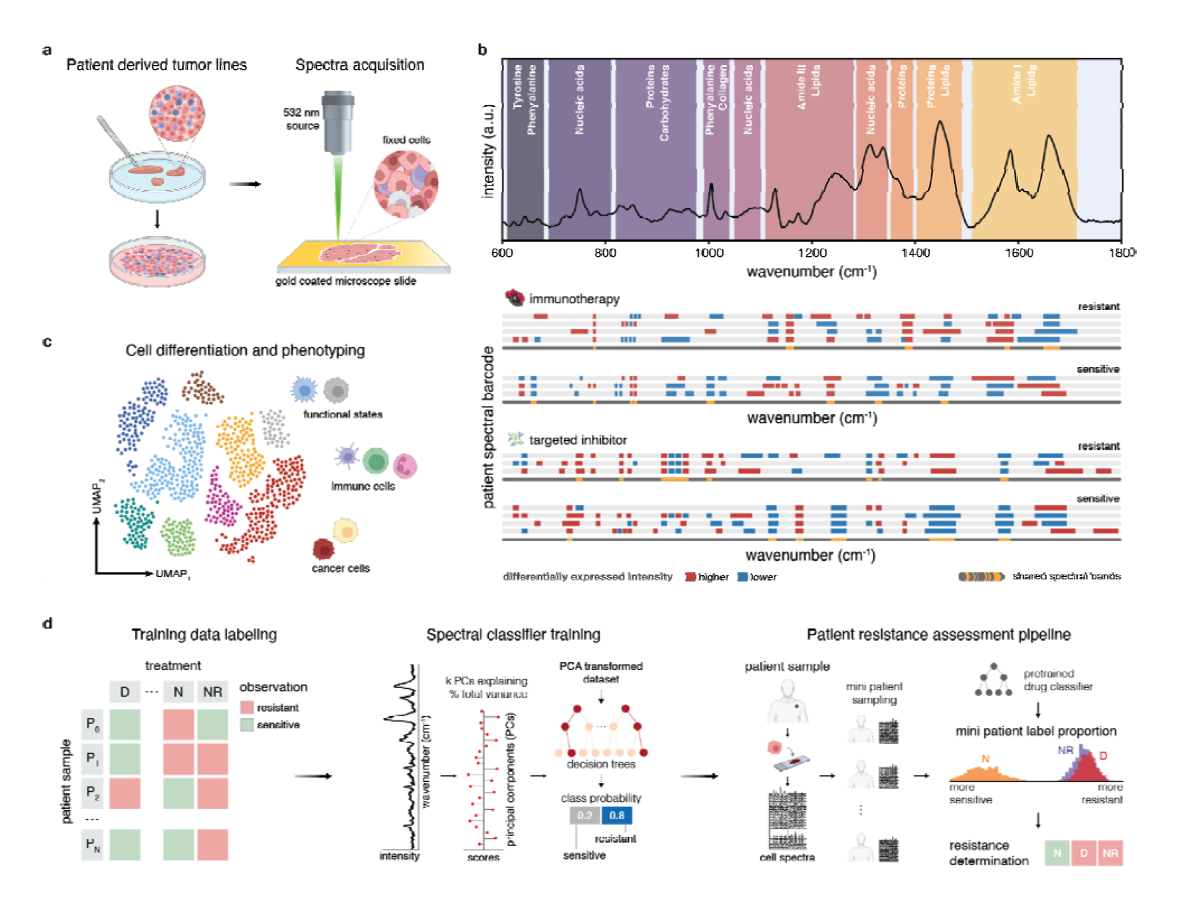
METHODS. We analyzed single-cell Raman spectra of mouse and human melanoma cell lines alongside nine melanoma patient-derived samples with known resistance profiles to targeted and immunotherapeutic inhibitors bemcentinib, cabozantinib, dabrafenib, nivolumab, and a combination of nivolumab and relatlimab. We assessed cell phenotyping classification and treatment resistance using random forests and feature importance analysis. For patient samples, we constructed a two-stage evaluation workflow to determine clinical drug resistance through aggregated single-cell predictions and identified corresponding highly variant spectral signatures using computational methods adapted from single-cell RNA sequencing methods.
RESULTS. In cell lines, our approach achieved >96% differentiation accuracy across tumor microenvironment cell types and induced functional phenotypes. Persistent (drug-resistant) cells formed subclusters based on genetic mutations rather than sample origin, with Raman signatures reflecting biochemical changes relevant to therapeutic pathways. For patient samples, our workflow correctly inferred resistance likelihoods for 30 of 33 clinically-relevant patient-drug combinations (91% accuracy).
CONCLUSION. Single-cell Raman spectroscopy combined with machine learning offers a scalable, prognostic platform to predict therapeutic resistance likelihood, with further potential to advance clinical, multi-omic biomarker efforts for melanoma. Our approach may improve firstand second-line therapy selection assessments for precision medicine by providing rapid, nondestructive prediction of therapeutic response based on cellular spectral profiles.